Services

Remote Analysis Desktops - BAND and BARD
Bioimage ANalysis Desktop (BAND) and Bioimage Analysis Research Desktop (BARD) are platforms that offer cloud-based desktops accessible with a web browser. So you can comfortably analyze your bioimage data with high computing power from anywhere with an internet connection.
Who are BAND and BARD recommended for?
- Life scientists
- Bioimage analysts
- Course organisers/trainers
Which infrastructure do you need?
- For BAND and BARD users: A workstation or laptop with internet connection
- For BAND deployment (e.g. system administrators, software developers): OpenStack or virtual machines
- For BARD deployment: Kubernetes clusters
Which expertise do you need?
What are BAND and BARD and how can you benefit from it?
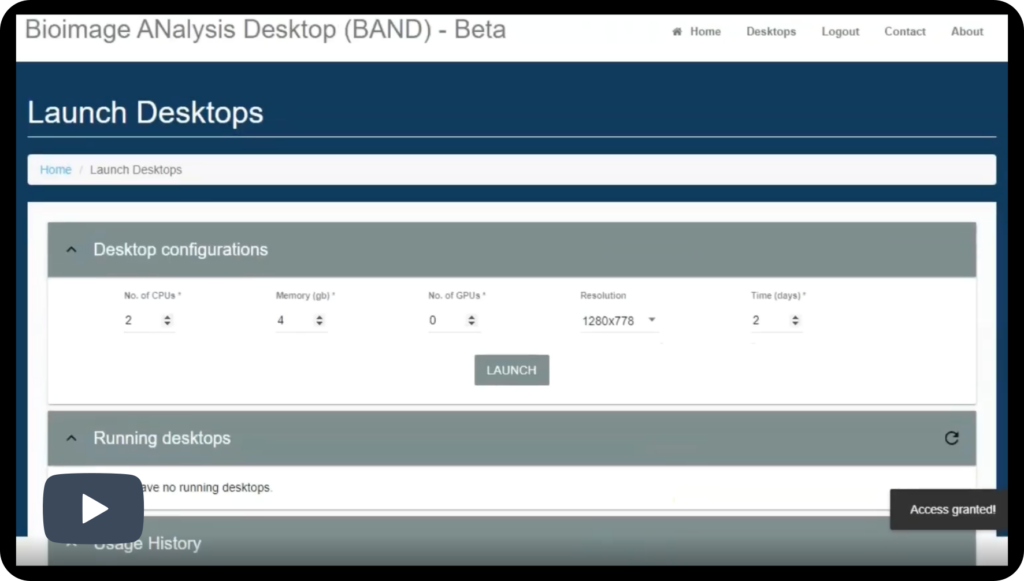
BAND is a cloud-based, flexible computing environment designed to make bioimage analysis easier and more accessible for users. It offers you essential tools and computing power in a single, integrated workspace. You work within a familiar graphical desktop that opens in a web browser. You will not need physical hardware, elaborate local installations or complex environment setup.
BAND is particularly suitable for your bioimage analysis workflows that require access to cluster computing. It is ideal for researchers who need both interactive exploration and large-scale batch analysis.
Users can launch desktops with resources tailored to their specific tasks, such as the required number of CPUs, memory, or GPU acceleration.
From a technical perspective, BAND is built on OpenStack, open source cloud computing infrastructure, and uses containerized software to provide a consistent and reproducible computing environment. Each user desktop runs as a cluster job with resources allocated on demand. Access to BAND is provided through federated login mechanisms, such as Google, LSAAI, EGI-Check which allow users to sign in securely using existing credentials.

How to get started:
- FOR ACCESS TO BARD: Request an account for access to the EMBL BARD instance by sending an email to cbbcssupport@embl.de.
- Once you have your account, open a browser and access the website of the instance.
- Select a ‚flavour‘ for your desktop by clicking the option buttons under “Resources”.
- Click ‚Connect with Guest Account‘ and enter your login details. Once loaded, you will see a desktop interface, your new workspace. The application dock is located at the bottom of the screen.
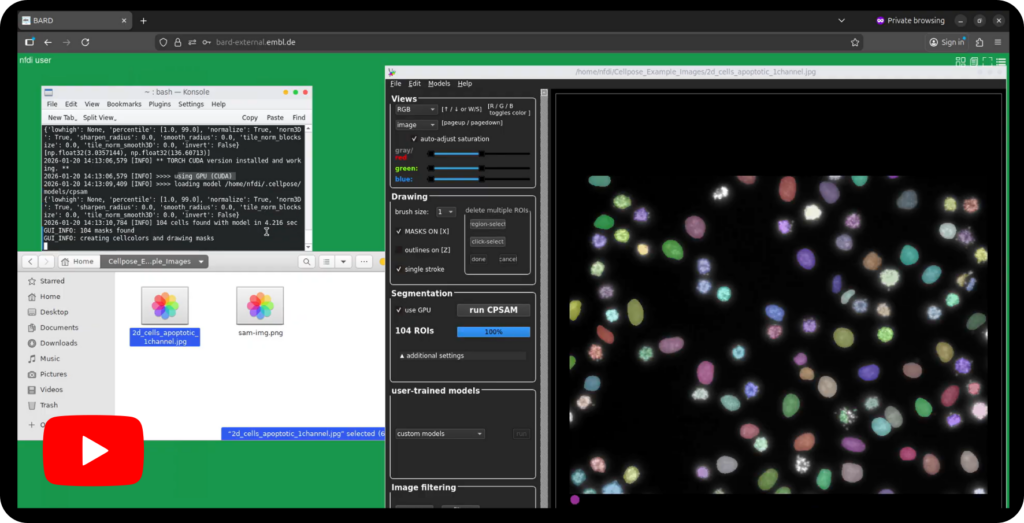
- Start your application of choice by clicking its icon in the bottom dock (for example Cellpose-SAM as in our video tutorial example). Off you are for your analysis. Additional applications can be accessed from the menu in the top-right corner.
- FOR ACESS TO BAND:
This part is still in progress, stay tuned for the update! Contact our Help Desk for urgent requests.