Services
Remote Analysis Desktops
This service allows you to analyze your microscopy data using powerful computing resources – anywhere, without any installations required. This so-called Desktop as a Service (DaaS) provides you access to computing resources such as CPUs and GPUs to run the analysis workflows your imaging project requires. All you need is an internet connection. NFDI4BIOIMAGE actively contributes and offers you 2 services for this: Bioimage ANalysis Desktop (BAND/BARD) and the Open Source Virtual Desktop Infrastructure (OSVDI). Explore and employ them.
Bioimage ANalysis Desktops (BAND/BARD)
BAND (Bioimage ANalysis Desktop) and BARD (Bioimage Analysis Research Desktop) are cloud-based desktops that empower you to analyse your microscopy images with with free access to powerful soft- and hardware via a browser. While BAND is accessible to everyone, BARD is is a service offered via collaborations with the EMBL. BAND provides you a ready-to-use image analysis environment including access to state-of-the-art community-developed tools. These include general software such as Fiji, napari, CellProfiler and KNIME as well as packages and tools for more specialized applications. Explore your options.
How can you access BAND to start your powerful image analysis?
FIRST TIME USERS of BAND, welcome. Initially and once, setup a login and register for access to remote desktops:
a) Login to LS Login: In case your local university or institution does not yet provide you access to it, register yourself first using e.g. your ORCID or GitHub logins (duration about 10 minutes). Once you can login to LS Login, you are prepared to create an account with BAND.
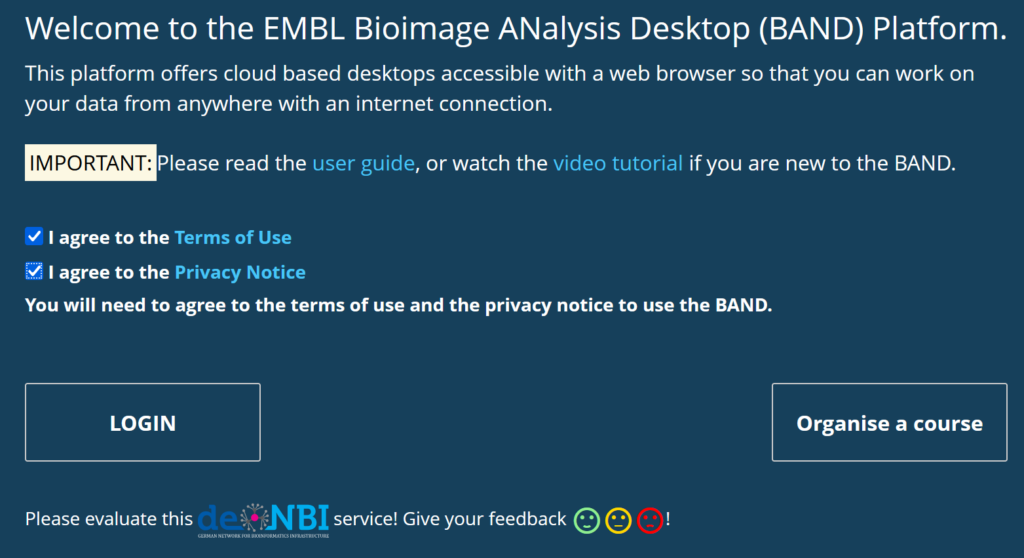
b) Create an account with BAND: Access the DeNBI BAND service in your browser. You MUST check to agree to the Terms of Use and the Privacy Notice buttons to be allowed to login. (In case your browser only shows 1 checkbox, change to a different web browser.)
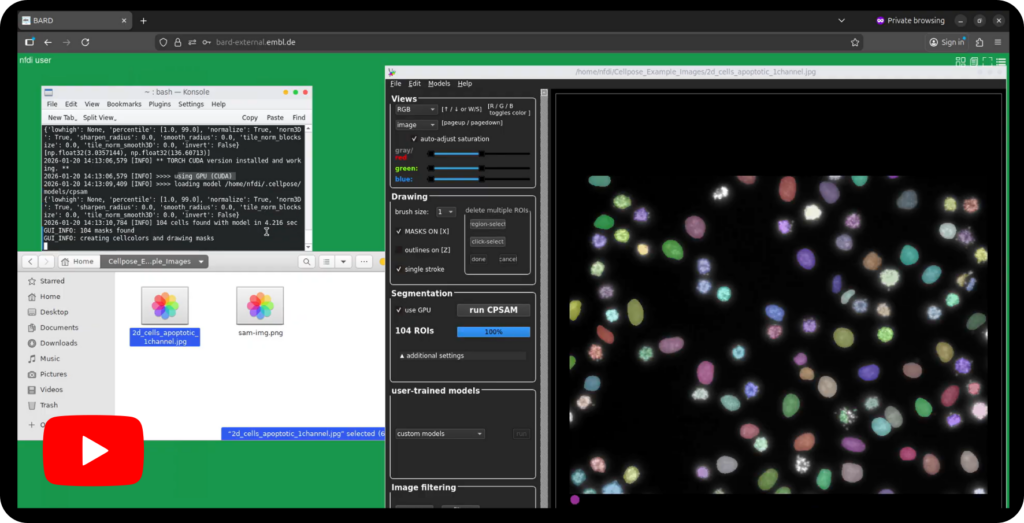
As registered user: Access BAND via LS Login and you arrive at the start page of BAND. To start a new desktop session, define the settings of the analysis desktop you aim to use by selecting required hardware components such as CPUs, GPUs and storage space and get access to a desktop tailored to your needs. In case you have a running desktop session, you can re-access it from the start page of BAND and continue where you had left.
Open Source Virtual Desktop Infrastructure (OSVDI)
The Open Source Virtual Desktop Infrastructure (OSVDI) we develop addresses the hardware accelerated full virtualization of traditional graphics workstations deployed in all aspects of interactive imaging. Test our demo hosted at the Computation Center of Universität Freiburg.
Who is OSVDI recommended for?
- IT infrastructure operators
- Operators of imaging facilities and labs
- Course organisers/ trainers and life scientists as users
Which expertise do you need?
- Users: A workstation or laptop with internet connection in a local of your choice
- IT/ facility or labs operators: resources for hardware and virtualization (not much different from ‘standard’ hardware in conventional setups)
Which infrastructure do you need?
- For OSVDI usage: no training required
- For image analysis via OSVDI: knowledge in bioimage analysis and the used applications
How to get started with OSVDI
- We will soon provide details to access the Freiburg demo instance of OSVDI. Stay tuned.
What is OSVDI and how can you benefit from it?
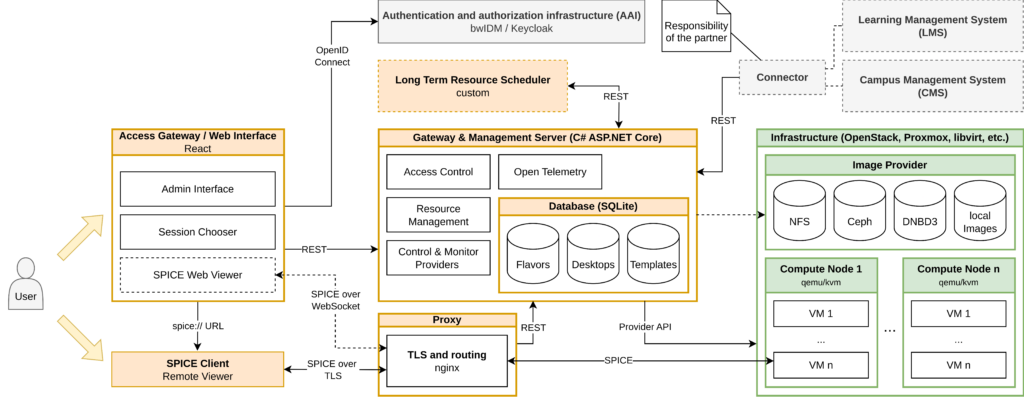
Virtual Desktop Infrastructure (VDI) solutions provide complete remote desktop representation in virtualization or cloud to be accessed over a network connection. OSVDI provides a full stack virtualization of (powerful) imaging workstations running a wide range of desktop operating systems and applications.
VDIs provide infrastructure and tools that provide researchers remote access to centralized hard- and software setups via virtual desktops with similar user experience to the local workstation and the known software environment. The desktop environment is identical to the traditional workstation. Even very high resolution displays are possible in case of appropriate network connection. VDI usage requires much lower network bandwidth for transfer of large datasets, ranging from10 Mbit/s in standard full-HD to less than 100 Mbit/s for significantly higher resolution and fast changes.
All VDI/DaaS solutions avoid the necessity for you as a user to actually come to the office to work on specialized hardware but rather handle and analyse data through remote access. Compared to traditional desktop computer environments, such centralized, virtualised and scalable cloud-like infrastructures are charactised by:
- Less administrative work
- Remote and faster data access
- Better access control
- Higher security
- More efficient and flexible hardware and software utilization
- Lower environmental footprint due to more efficient resource usage
OSVDI is co-funded by NFDI4BIOIMAGE and the bwCloud3 service development project.