Wall Calendar 2026
Our third edition of the NFDI4BIOIMAGE calendar highlights bioimage repositories
This year, we turn our attention to the “end” of bioimage data — the moment of publication. While publishing might seem like the final step in the research process, the point where a project comes to completion, we all know it is far from the end. In reality, publication marks the beginning of new rounds of discoveries: once shared, bioimage datasets can be reused, reanalyzed, and reinterpreted by other researchers.
To highlight this crucial stage in the data lifecycle, the 2026 edition of our calendar focuses on repositories for publishing bioimage datasets. Each month presents one exemplary data publication and introduces its hosting repository. Together, they illustrate the diversity of platforms available for storing, sharing, and preserving bioimage data in a way that is sustainable and FAIR — Findable, Accessible, Interoperable, and Reusable. By scanning the QR code on each calendar page, you can explore an accompanying webpage that provides direct access to the published dataset, detailed information about the data, and insights into the respective repository.
We teamed up with Henning Falk, a scientist by training who also creates cartoons for the bioimaging community. Henning helped us visualize the efforts of the NFDI4BIOIMAGE consortium and its team of data stewards to make bioimage data FAIR, turning complex processes into clear and engaging illustrations.
Unlike in previous years, where “The Beauty of Metadata” was visualized directly on the calendar pages, this year’s focus on published datasets means that all biological metadata can now be found within the corresponding repositories. Metadata form a necessary and indispensable part of every data publication — ensuring that datasets remain understandable, searchable, and truly reusable for the scientific community.
With this year’s calendar, we invite you to explore how data publication transforms research results into shared scientific resources, creating new connections, fostering collaboration, and ensuring that bioimage data continue to inspire discovery long after their initial generation.
How to contact us?
For help or advice on your bioimage data don’t hesitate to reach out to the NFDI4BIOIMAGE Data Stewardship Team via our help desk request form or by email.
For further information you can also reach out to our project office or check out our webpage.
Funding Information
This calendar is published by the Heinrich Heine University Düsseldorf as a contribution to the project NFDI4BIOIMAGE, funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under the National Research Data Infrastructure – NFDI46/1 – 501864659.
The NFDI4BIOIMAGE consortium consists of legally independent partners and does not act autonomously towards third parties. All institutions represent their respective contributions and work together for the course of the project.
January 2026: BioImage Archive – a repository for all kinds of imaging data
By our data steward Ksenia Krooß
If you are on the verge of publishing your paper, you should consider publishing the corresponding data along with it. While some journals or funders might request this, in most cases it is up to you to decide whether to publish your data. Of course, publishing data takes some effort, but by doing so you provide a great service to the entire scientific community. By sharing your data, you make it more FAIR, which stands for Findable, Accessible, Interoperable, and Reusable. FAIR data benefits other researchers, as they can save time and resources if relevant studies already exist and the data and metadata are accessible for reuse.
One repository that can serve this purpose is the BioImage Archive (BIA). This repository is hosted at the European Molecular Biology Laboratory – European Bioinformatics Institute (EMBL-EBI) in Hinxton, UK, and has a steadily growing portfolio of studies covering a plethora of organisms and microscopy techniques (as of January 2025: 600 TB across more than 800 studies). Since it is not just about your data but also the accompanying metadata, you can provide extensive metadata in the BIA submission tool and through the File Lists that accompany each of your experimental items (the so-called Study Components). The metadata schema is based on Recommended Metadata for Biological Images (REMBI), a widely used guideline within the bioimaging community. After publishing your work on BIA, your study will be findable and accessible, and most importantly, reusable, as researchers can download individual Study Components or the entire study for their own research purposes.
Preparing your data for data publication can feel overwhelming when you approach this process for the first time. Often researchers are not sure where to publish their data or feel uncertain about all the new terminology introduced by the metadata schema in the submission process. If that sounds familiar, you can reach out to our NFDI4BIOIMAGE Helpdesk for support. A short email describing your bioimage data management struggle is enough to get in contact with us. As a first step, one of our Data Stewards will contact you. Your request might be resolved quickly by pointing you to the right tool or person, or, if it involves data publication, the Data Steward will schedule a meeting with you (online or in person).
In this first meeting, we try to understand your study: What organisms and tissue did you use? What microscopy techniques were involved? Did you run any analyses? Do you have only raw data, or also intermediate or processed results? This information helps us recommend the most suitable bioimage repository for your data. For instance, BIA does not accept patient data but any other bioimaging data in general. Is your data more centered around cells and tissue and novel imaging techniques, then the Image Data Resource (IDR) might be a better fit.
After identifying the right repository, we discuss its specific requirements. In the case of BIA, we try to point out Study Components within your study. As mentioned earlier, these are experimental items in your study and can stand alone as an independent entity (what is a Study Component). Each Study Component consists of several modules based on the aforementioned REMBI framework. We guide you through these modules in the submission tool and explain what metadata is needed (see the submission guide of BIA).
Often, this concludes our first session, and we leave you with your homework: to organize your data according to the identified Study Components and collect the necessary metadata for each module. In a follow-up meeting (or sometimes already during the first), we evaluate the data organization once again and discuss the File List that accompanies every Study Component (what is a File List). These File Lists list all files within a Study Component and include corresponding metadata. The metadata in the File List focuses on the differences among files to make your data even more understandable and is a mandatory part of the data publication. When other researchers download one or more Study Components from your study, they receive these File Lists along with your data, ensuring they have complete and well-documented datasets.
We also cover how to upload your files to the submission tool, discuss the available options depending on your data size, and of course assist you if you encounter any issues. Once your files are uploaded, your File Lists complete, and all modules filled out in the submission tool, you’ll be ready to publish your data. Conveniently, you can set a preferred release date, allowing you to coordinate the data publication with the release of your paper.
The author of this month’s image, Sarah Knapp (RWTH Aachen University, University Hospital Aachen), is a great example of this established workflow. She contacted us via our Help Desk after learning about our services during a symposium. In our first meeting, we discussed how to structure her data and went through the entire submission process together. After the meeting, I sent her some additional information and files to help her prepare for the data publication. We stayed in touch via email, and I checked on her progress. Sarah managed to prepare all her data accordingly in a short amount of time. She encountered some issues during the file upload but solved them herself and even shared the solution with me, which is very valuable information for future requests. I was very pleased with the outcome and happy to see her data successfully published.
As you can see, publishing your data involves several steps, but you do not have to do it alone. Reach out to the NFDI4BIOIMAGE Helpdesk and get the support you need to successfully, and as FAIR as possible, publish your data.
February 2026: OMERO – a bioimaging database with the option for controlled “open” access
By the image author Nadja Rotte and our data steward Jens Wendt
The image originates from our research on the genetic causes of male infertility, including the description and functional characterisation of the human M1AP gene, which plays a crucial role in meiotic recombination. In our studies, we described pathogenic variants in M1AP as a recurrent cause of non-obstructive azoospermia (absence of spermatozoa in the ejaculate) and cryptozoospermia (few spermatozoa detected after centrifugation). These men typically exhibit a germ cell arrest at the spermatocyte stage, reflecting a failure of spermatogenesis at the level of meiosis.
Interestingly, a small subset of men with M1AP-associated infertility still develop rare round and elongated spermatids, indicating that partial meiotic progression can occur and haploid germ cells can develop. Identifying such residual spermatogenesis is clinically relevant, as it may open the possibility of testicular sperm retrieval (TESE) and medically assisted reproduction to enable fatherhood. Beyond its diagnostic implications, studying M1AP also provides critical insight into the molecular regulation of meiosis and helps to refine gene panels used for genetic testing in men with unexplained infertility and spermatogenic failure.
At the Institute of Reproductive Genetics in Münster, testicular biopsy samples from men who consent to participate in research are analysed using immunohistochemical staining to visualise specific germ cell markers such as CREM. These biopsies originate from TESE procedures performed at the Centre of Reproductive Medicine and Andrology in Münster and at the Clinic and Polyclinic for Urology in Gießen. Combining these histopathological analyses with a whole-slide microscopic scanning system results in comprehensive, high-resolution assessments of entire testicular sections even years later. Besides, re-analysis and digital archiving of valuable human tissue samples becomes possible.
In collaboration with the Imaging Network core facility of our university and NFDI4BIOIMAGE, these digital data have been processed and integrated into OMERO (Open Microscopy Environment Remote Object), a secure and privacy-compliant image data platform. This not only preserves the histological information in a sustainable format but also enables controlled access for other researchers. Rather than relying on limited image snapshots in publications, researcher can now explore our complete human testis sections – an approach that promotes transparency, reproducibility, and interdisciplinary use of these unique research materials.
Unfortunately, this imaging data was initially captured in a proprietary file format accessible only through the manufacturer’s viewer software. The export from there generated Big TIFF files containing three separate images: a whole slide overview, the slide label, and the actual high-resolution slide scan. To streamline the dataset and convert it into an open, interoperable format, the NGFF converter tool was utilized. Its user-friendly graphical interface allowed for easy selection of the specific image series needed—in this case, the primary slide scan—while excluding the redundant overview and label images. The tool seamlessly converted the selected series into OME-TIFF format, ensuring the data adheres to open microscopy standards and is accessible for broad reuse in the scientific community without dependency on proprietary software.
Ultimately, our work on M1AP exemplifies a broader effort to bridge clinical diagnostics and molecular biology in male infertility research. By linking genetic findings to histological and cellular phenotypes, we aim to establish a more precise genotype-phenotype correlation, identify new infertility genes, and improve diagnostic and therapeutic options for affected men and couples.
March 2026: Image Data Resource – the Original
By our data steward Jens Wendt
When it comes to publishing bioimaging data, choosing the right repository is a crucial decision. While the BioImage Archive (introduced in our January edition) accepts a broad range of imaging data, some datasets benefit from being hosted in a more specialized environment — one that not only stores the data but also makes it directly viewable, queryable, and interactable for the scientific community. The Image Data Resource (IDR) is exactly such a platform.
The IDR is a public repository of reference image datasets, hosted at the European Molecular Biology Laboratory – European Bioinformatics Institute (EMBL-EBI) in collaboration with the University of Dundee. Unlike general-purpose archives, IDR focuses on curated, high-quality datasets that serve as reference resources for the community. A key feature of IDR is that it is based on OMERO (which we introduced in February), providing a powerful web-based viewer that allows researchers to browse and explore published image data directly in their browser. Each dataset in IDR is richly annotated with biological and experimental metadata, and the repository links imaging data to other public resources such as genes, phenotypes, and chemical compounds. This tight integration of images with structured, searchable metadata makes IDR a particularly valuable resource for researchers looking to re-examine published image data in new contexts or to validate their own findings against existing reference datasets.
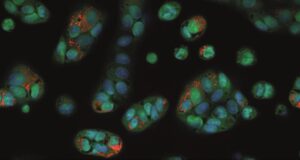
This month’s image originates from a large-scale interdisciplinary study on drug-induced phospholipidosis (PLD), a collaboration between the Fraunhofer Institute for Translational Medicine and Pharmacology (ITMP) in Hamburg and the Karolinska Institutet in Stockholm. The study investigated more than 5,000 compounds from a drug repurposing collection for their propensity to induce PLD in different cell lines, and developed machine learning models to predict this cellular adverse effect. The imaging data generated in this study — showing Vero-E6 cells with markers of phospholipidosis — was published on IDR, making the complete high-content screening images accessible and reusable for the wider research community.
But what exactly is phospholipidosis? PLD is a storage disorder characterised by the excessive accumulation of phospholipids within lysosomes, often induced by certain drugs. This phenomenon is particularly relevant in drug discovery because PLD can confound the results of cell-based screens — for example, during the SARS-CoV-2 pandemic, some compounds initially identified as promising antiviral candidates turned out to be PLD inducers whose apparent activity could not be replicated in vivo. The image on this month’s calendar page shows Vero-E6 cells treated with such a PLD-inducing compound, with nuclei stained in blue, cytoplasm in red, and accumulated phospholipids visible as bright orange spots.
This publication was the first instance where an NFDI4BIOIMAGE Data Steward cooperated with IDR and took over a significant portion of the upload and annotation process. First of, a publication in IDR requires a prior submission to the BioImage Archive, as IDR sources its data from BIA — an integration that highlights how closely these two repositories already work together and that is set to deepen further spearheaded by the foundingGIDE initiative. Additionally, the imaging data needed to be provided in the OME-Zarr format, a cloud-optimized, chunked file format that is rapidly becoming the new standard for bioimaging data and enables the seamless browser-based viewing experience that IDR offers through OMERO. Beyond the file format, much emphasis was placed on linking the experimental metadata to established ontology terms, ensuring that the dataset is not only human-readable but also machine-interpretable and truly interoperable. Overall, both the data upload and the metadata annotation required a considerable level of technical knowledge — from handling large-scale file conversions to navigating bioimage-specific ontologies — making this a process where support from a data steward can make a real difference.
Publishing imaging data from large screening campaigns in a repository like IDR goes beyond simply fulfilling a data sharing requirement. It transforms raw screening results into a community resource. Other researchers can explore the images, verify reported findings, and build upon the dataset for their own studies — whether that means training new machine learning models, benchmarking image analysis pipelines, or investigating the cellular phenotypes associated with specific compound classes. This is an integral part of FAIR data: making research outputs not just available, but truly reusable.
Data accessible on IDR: publication is pending – stay tuned
Data accessible on BIA: https://www.ebi.ac.uk/biostudies/bioimages/studies/S-BIAD2282
April 2026: FAIR Instead of Lost: How Data Stewardship Makes Imaging Data Sustainable
By Isabel Kemmer, FAIR Image Data Steward at Euro-BioImaging
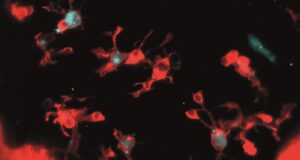
This month’s image is a SPIM image of the developing zebrafish brain displaying microglia, immune cells responsible for clearing dying neurons. This image in particular reflects an experiment where alongside normal single-centrosome microglia, a population modified to possess two centrosomes is visible. These double-centrosome microglia demonstrate increased phagocytic efficiency, linking centrosome organization to the rate of neuronal clearance.
This image is part of a much larger dataset on zebrafish brain microglia, available in the BioImage Archive. The original author and depositor of the dataset, Katrín Möller, University of Iceland, recalls: “after curation, like maximum projections etc., it was still 650 GB of data. And yes, my institute had servers […] however they couldn’t store all this data forever and ever and I found it a pity that my 5 years of imaging work would just be deleted at some point. So, I was really desperately looking for something to do with my dataset.”
Her experience reflects a challenge faced by many researchers: what happens to large imaging datasets once a project ends? And who has the time and takes responsibility to take care of them?
That’s where Isabel Kemmer, FAIR Image Data Steward at Euro-BioImaging, got involved and together, they deposited the dataset in the BioImage Archive. This process was carefully guided to ensure that the data was curated and stored according to FAIR principles, making it Findable, Accessible, Interoperable, and Reusable. This support was provided through Euro-BioImaging’s Data Stewardship Service, launched in 2022, which provides guidance for staff and researchers in managing and depositing imaging data. As such, data stewardship is one element of the distributed research infrastructure Euro-BioImaging which provides open access to biological and biomedical imaging instruments, training, and data services across Europe.
“I had just launched this service when my colleague Johanna came back from the BNMI Symposium and told me about a researcher looking for help with data deposition. When I spoke with Katrín and saw her striking images full of scientific value, I knew this was the perfect start,” reflects Isabel on her experience, “My process of supporting researchers with data deposition has evolved since then and become more streamlined, but at its core it remains the same.
I begin by understanding the project and grouping the study into REMBI study components. Sometimes this is straightforward, while other times, we discuss options with the researcher to find the most logical order that best represents the study. This outside perspective lets me spot gaps in data organization where our views don’t align, which then can be addressed. Once the study component framework is set, the rest of the REMBI metadata falls into place.
I often provide prefilled metadata templates, as I did with Katrín, or suggest which general metadata should be recorded based on early discussions. We try to incorporate ontologies, when possible, where I provide guidance to find appropriate terms. Meanwhile, the depositors might reorganize their data to better match the study components if they are not already aligned.
After the data is uploaded to the BioImage Archive it’s not yet time to hit the submit button. Instead, the final crucial step is generating and refining the file list, which is a table that captures differences between files within the same study component. In this way, each image gets assigned unique metadata which can’t be described by the attributes of the REMBI study components alone. A well-curated file list is usually what makes entries stand out.”
Katrín also highlights the importance of metadata curation: “The better you annotate the images, the better it is for the next user or whoever wants to explore your data because if you put very few annotations people don’t really know what to do with it. […] In my case, the most important metadata were the zebrafish lines and organelle markers, and the spatial and temporal image resolution.”
“I always say ‘start thinking about data deposition immediately, ideally before or while you are collecting your data’. If you already properly organise your files and link relevant metadata from the beginning, it makes deposition much easier down the line.” concludes Isabel.
While early planning makes the process of deposing smoother, sharing well-curated data always opens the door to new possibilities, as Katrín’s experience shows: “I can now also share my data with the world easily. […] If a scientist uses this dataset first, before acquiring their own data, it could potentially save them a step. It could give them a starting point to test their pipelines, or allow them to compare behavior of zebrafish microglia to another system. […] I would of course be very excited if we could learn something new about microglia using my data”.
Beyond being able to share data, Katríns dataset is also featured in BioImage Archive’s visual gallery which showcases selected datasets in the cloud-optimised next-generation file format OME-Zarr. This way, the data can now also be explored in-browser and easily analysed in the cloud. Sharing data in a way that others can reuse it easily not only speeds up new discoveries but also helps make science more reliable.
“Despite the hard work, I’m really glad I did this. It feels like a big step towards more sustainable science,” concludes Katrín. “Afterall, the more of your data you share, the more believable your study is”.
Full dataset in BioImage Archive: https://www.ebi.ac.uk/biostudies/BioImages/studies/S-BIAD564
Dataset in OME-Zarr format: https://www.ebi.ac.uk/bioimage-archive/galleries/S-BIAD564.html
News article about the release of the dataset: https://www.eurobioimaging.eu/news/neuroscience-new-microglia-dataset-available-on-bioimage-archive/
Video featuring Katrín and Isabel talking about this dataset: https://www.youtube.com/watch?v=-1wg0LDCBco
Original publication: https://doi.org/10.7554/eLife.82094

May 2026:
Coming in May, stay tuned!

June 2026:
Coming in June, stay tuned!

July 2026:

August 2026:

September 2026:

October 2026:
Coming in October, stay tuned!

November 2026: