Services

Format Development and Ambassadorship
NFDI4BIOIMAGE research software engineers actively serve the community through development and implementation of next generation file formats, so called NGFFs. The most prominent example, OME-Zarr, is aimed to contain multidimensional microscopy datasets of as diverse dimensionality and microscope techniques and system as possible combined with OME-standardized metadata. This development transforms the plethora of data formats towards community-driven standardization to simplify data storage and analysis and harmonizes data handling overall. The consortium with its data stewardship team supports this development through bridging of communities and advocacy of the implementation across tools and institutes.
What are next generation file formats and how can you benefit from them?
Over the years, microscopy data has become larger in size and complexity and the diversity of microscopic systems from diverse vendors has created a multitude of bioimage file formats. Traditional file formats no longer fully meet the needs to manage modern bioimage datasets in an easy, fast and efficient way. This has motivated the mission to specify and bring into daily research life next-generation file formats in agreement with both technical standards and user needs. NFDI4BIOIMAGE actively contributes to this development and fosters exchange of stake holders in this process.
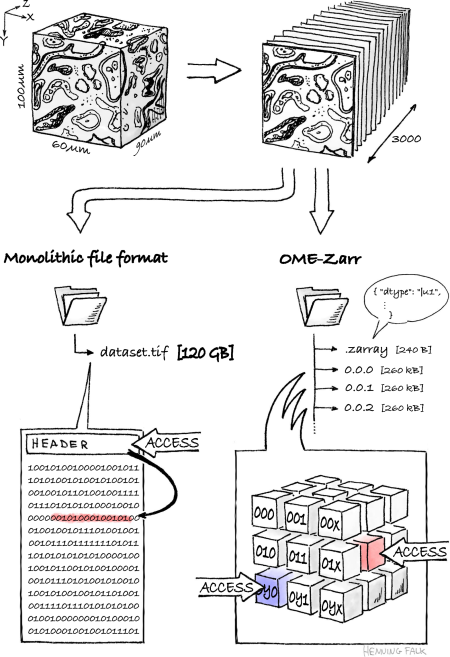
Next-generation file formats such as the format Zarr are chopped into a multitude of tiny blocks termed chunks that are each independently accessible. This architectural change from traditional file formats being mostly a single monolithic block of data to next-generation file formats allows reading and writing of only those data chunks of interest. Scientists can must faster visualise or analyse the image dataset part of interest, being for example specific substructures of the investigated sample.
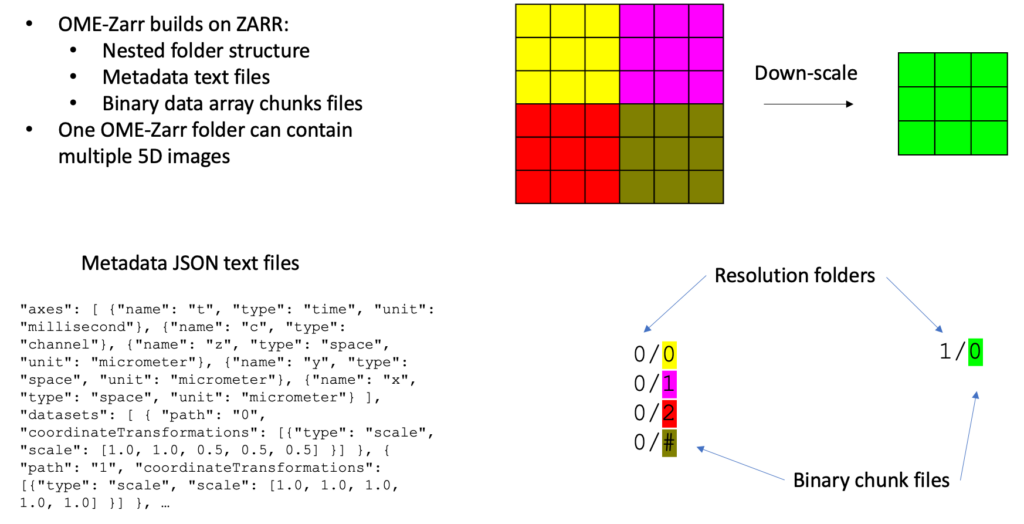
Bioimage data in the Zarr format is accompanied by rich metadata in the community-developed Research Object Crate (RO-Crate) format implementing the Open Microscopy Environment (OME) data model to use precisely defined terminology. This development further launched and drives the usage of web-based viewers for bioimage datasets such as vizarr and Neuroglancer amongst others. NFDI4BIOIMAGE research software engineers ensure the implementation of OME-Zarr into state-of-the-art tools and services developed in the consortium.
Learn more about OME-Zarr and the currently ongoing process to define and advance its specifications to best serve the community. Or get a first impression of the OME-Zarr data structure and accompanying metadata files here.
Can you contribute to the development of next generation file formats?
Yes, you are warmly invited to contribute. Learn how you can do this or reach out to our Help Desk for support.